During this past week, Martín and I had the pleasure of attending Epihack Tanzania 2014 in Arusha as InSTEDD facilitators of the event. This Epihack, 3rd of its nature, was organised by SACIDS with the support of the Skoll Global Threats Fund.

Epihack Tanzania 2014 organised by SACIDS
The facilitator team was integrated by the two us, plus Channe Suy and Sokmesa Khiev from iLab SEA, as well as Keng Susumpow and Bunny Petra from Opendream Thailand, who shared their experiences from previous events in Chiang Mai (Thailand) and Laos. Professors Esron Karimuribo, Andrew Kitua and Mark Rweyemamu, and Eric Beda and Mpoki Mwabukusi from the SACIDS team further integrated the team, which was completed by Mark Smolinski, Jennifer Olsen and Kraig Butru from Skoll Global Threats Fund.
The goal of this Epihack was to bring together human and animal health experts with developers, in order to design and prototype solutions aimed at improving outbreaks detection using a One Health approach; this concept recognises that the health of humans is connected to the health of animals and the environment, and thus seeks to share and combine information between these two sources.
The event was truly fantastic, as it brought together awesome people with endless energy who engaged in the most interesting discussions and solutions; and the results were no less impressive. But let’s do a quick recap of the event day-by-day first.
Day 1: Exploring the problem space
The first day, after the mandatory introductions and presentations, found ourselves with a vast problem space to be explored. Though improving outbreak detection was our ultimate goal, disease surveillance has multiple aspects to it, so the first task was to be able to identify the challenges that affected the health experts. We split into random groups, as heterogeneous as possible, combining people with different backgrounds and from different countries, and started a free generative process to navigate the problems that affected the experts.

Martin presenting the findings after the first brainstorming
After some very interesting explanations from the experts, each group presented a set of the most pressing problems, which we were able to narrow down to five main areas by combining the issues that were heard across all groups:
- Community participatory surveillance
- Official data reporting channels
- Feedback and 2-way communication
- Contact tracing
- Human and animal data integration
Having identified these areas, groups were rearranged by having everyone choose the problem that was most interesting to him or her, ensuring a good balance between health experts and developers; and proceeded to delve deeper into the issues that affected each area after assembling each team.
Day 2: Field trip
The second day was a very particular one, as we visited Ngorongoro, which turned out to be one of the most beautiful places I’ve ever seen.
Having a 3-hour trip to Ngorongoro, we split in vans according to our groups, and started the process of narrowing down the problems identified on the previous day in order to start designing a solution. We then visited local health facilities, Maasai communities, and exchanged views with local community health workers. As it is so often said at InSTEDD, if you don’t go, you don’t know.
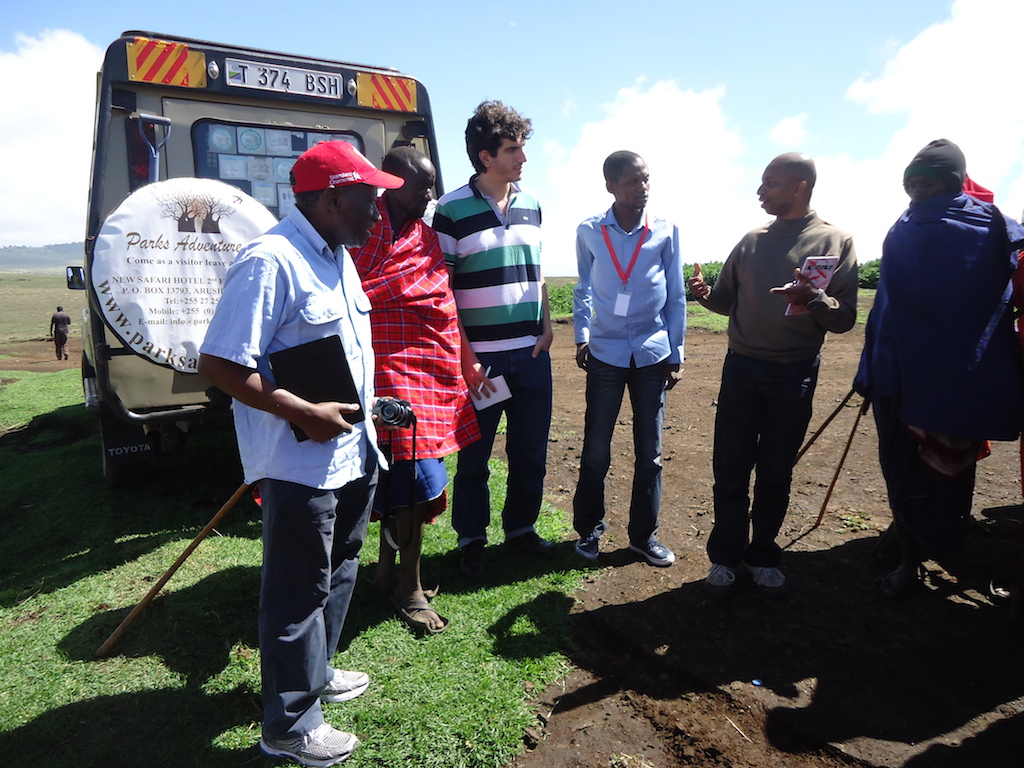

Visiting a Masai village in Ngorongoro
This allowed us to obtain vital information on the reality of disease detection in the field, which provided a unique opportunity to validate our previous assumptions and pivot as necessary to design a solution tailored to the real local needs.
We were able to see that smartphone penetration was far from complete, and though community health workers could be equipped with one, having entire villages without electricity made frequent battery recharging impossible. 3G network coverage was also a problem; this led us to look for simpler solutions as we had done in the past.
After this validation session, we made a short visit to the Ngorongoro crater, a pristine beautiful area, rich in wildlife, with a variety of species I had never witnessed. Though out of the Epihack scope, this visit was a truly wonderful experience, and undoubtedly one of the highlights of the week.
Day 3: Start hacking
Having concretised the areas into particular problems, on the third day we completed the design of the solutions during the morning, shared our ideas with each other group, and after noon, started hacking.
Even though each group had been assigned a different area, it was remarkable to see multiple overlaps between them, as each group had individually unveiled the same critical points by approaching the disease detection problem from different angles. Feedback and data sharing would come up in all discussions, and these in turn were based on the reporting and tracing channels designed by other teams.
After having a clearly defined path, developers jumped into hacking in no time. From InSTEDD, we shared with them several platform tools to help them jumpstart the prototypes and avoid reinventing the wheel; tools such as mbuilder relying on the Android local gateway for rapid SMS prototyping, verboice for managing voice based channels, and resource map for resource tracking, were quickly adopted by several teams. This also was a great opportunity for us to see our tools in the hands of fellow developers as they approached them for the first time, providing invaluable feedback to further improve them and reduce their learning curve.
Day 4: Hacking non-stop
One of the most remarkable things from days three and four was the endless energy of the developers. It was possible to witness them working hard on their problems until late hours in the night, and at some points, even early in the morning. Their commitment to building high quality prototypes was exemplary, with many pulling all-nighters or catching a quick nap before going back to work the following morning.

Martín on a 6 to 8 AM nap before starting the fourth day
But the health experts were no exception to the enthusiasm floating in the air. While some of them with a knack for technology remained next to the developers learning on the implementation details and the tools involved in them, others assembled in new health-experts-only groups that discussed opportunities for data management and sharing, and compiled a set of guidelines and symptoms of diseases of interest, both animal and human.
Day 5: Presentations
The results of the efforts of a week of hacking were unveiled on a presentation on the morning of the fifth day. Except for the group focused on data mangement, which facilitated expert discussions during the week, each of the other groups presented the problem, the proposed solution and a working prototype.
These prototypes help in not only providing a concrete implementation of the designed solutions, but also provide a conduit for feedback from the experts who can directly engage with the tools, and validate the hypothesis that each group had on how their approach can lead to better disease detection.
Community surveillance
The community surveillance group, facilitated by Keng and Eric, was the first to present their work. Reporting at the grassroots level was key for early disease detection, so providing community workers with the tools and incentives to easily notify the health system of any observed symptoms was a key component of the overall goal.
The One Health disease tracking map, prototyped during one of the first Epihacks and further developed by Opendream, was chosen as the platform to store disease reports; its smartphone powered application was also used to collect information from the field.
As an added value, and to increase workers involvement through feedback, this group worked closely with the experts to set up an expert system which would identify the reported disease with a certain confidence based on the symptoms reported, and yield recommendations based on this. Since automatic identification carries a potentially high error rate, recommendations were such to ensure that no counter productive behaviour would be encouraged, and the community would not take in their own hands tasks that were required to be carried out by experts.
Having the freshly uploaded report, this prototype answered on the spot with a recommendation on how to act based on its symptoms, and also leveraged the feedback prototype (see below) by having an additional channel to send further recommendations to target sectors.
Feedback
The second group to showcase their work was the one working on Feedback and 2-way communication, which I had the pleasure to facilitate. Even before the Epihack started, experts from SACIDS had identified as an issue the lack of information shared back to the workers who reported incidents. Volunteers sending information up the chain on witnessed symptoms were kept out of the loop, and failed to see the enormous value that was being generated by their contributions.
A need to keep a feedback loop was easily found, in order to foster the engagement of volunteers and increase the communities’ trust in the system, and also to empower them with the contextual knowledge required to react to the very situations they are reporting.
A first solution was to design a system hooked to the official reporting workflow, which automatically sends notifications back to the initial reporter to keep him informed on the status of the report; but this solution required extensive analysis and unfortunately fell outside of what could be done in the context of the hackathon.
The implemented prototype, named mtoolbox, was a platform for district-level health experts to broadcast contextualised SMS notifications to health facility workers, community health workers and the very community as well. Health experts, after observing a rise in certain symptoms in an area, could issue an alert to the community health workers to be on the lookout for similar cases, and send recommendations to the community on how to act, thus also promoting behaviour change.
The added value of this prototype was a viral subscription mechanism, in which people could either subscribe to the system or subscribe someone else, by indicating their role and location. It was on purpose that no validation on these fields was imposed, in order to make the subscription as smooth as possible; so users of this tool could review and refine the contacts database as needed.
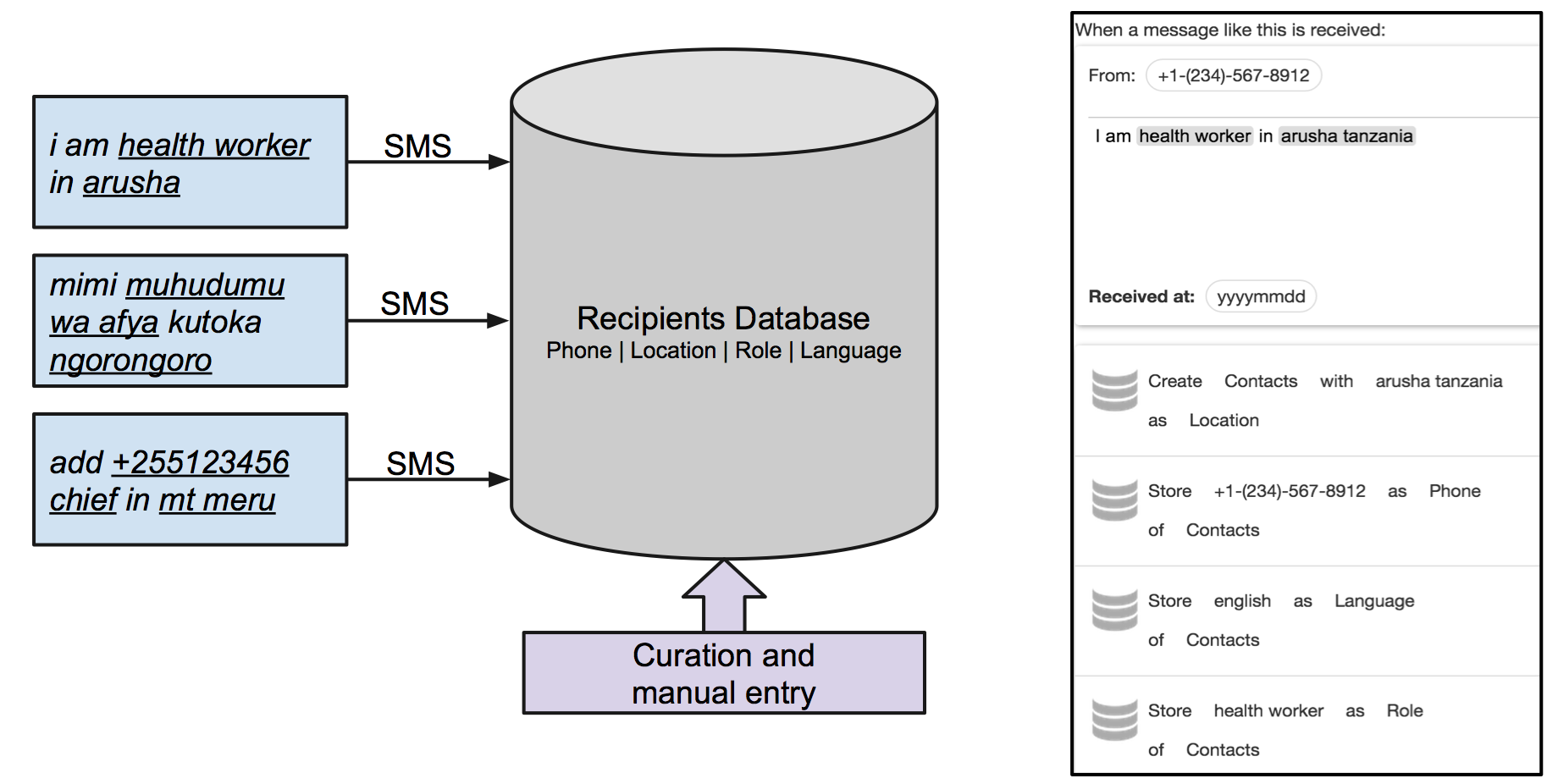
mtoolbox viral subscription
Both message broadcasting and the subscription flows were powered by mbuilder, and a client only webpage was set up to ease the process of sending messages to target sectors. A local gateway was set up to manage SMS messages, thus requiring a single Android phone to be the only hardware resource for the entire prototype.
Contact tracing
Working closely with Martin Simuunza from Zambia and Asad Islam from CDC, the group led by Martín presented their complete solution for animal contact tracing.
Livestock herds have particular points where they gather, such as water sources, pastures or markets, which are potential areas for contagion. This provides a unique opportunity to trace back the potentially affected animals when a case of an infectious disease is detected.

Dr Martin Simuunza presenting the motivation for the animal contact tracing app
By keeping a registry on these gathering points in resource map along with their catchment areas, any health officer can easily trace back which herds had contact with an affected animal in the last days. A Rails based website was quickly set up on Heroku for these operations, and also acted as a dashboard to keep track of reported events.
Such events were notifiable by farmers by placing a call to a verboice powered hotline, which (with recordings from Martin Simuunza) would ask for specific symptoms to provide the health expert observing the case with the required information to perform an initial analysis.
This hotline also acted as an entry point for the farmers to view the status of their previous reports, by making available the workflow information stored in the system, thus providing a first implementation of the workflow feedback design ellaborated on the previous group.
Official reporting
Last but not least, the group led by iLab SEA’s Channe showed their work on improving official reporting. Since internet access can be a challenge in several health facilities, SMS based reporting was sought. Three channels were set up, all of them converging to a single repository of data.
The first was structured SMS reporting. By uploading messages with a specified format, the reports sent by the health facilities could be handled by an mbuilder trigger, which parsed and stored the information uploaded.
Leveraging a tool shared by Moses Moine from Medic Mobile, a second channel was set up by using a phone application. Moses could design a small feature-phone application and burn it into a SIM using the appropriate technology, thus providing low-cost ubiquitous phones with a small app that could be used to generate structured reports, then transmitted via SMS in the background using a format opaque to the end user.
The third channel made use of the reporting wheel, which is a smartly designed paper-based artifact that provides numeric redundant codes based on a report that depends on a set of pre-defined categories. As well as the previous tool, it eliminates the complication of having to train staff on potentially difficult formats that need to be observed every time a report is sent.
Amazing gadget? This piece of paper: Reporting Wheel by @InSTEDD @EpiHackTanzania http://t.co/iFOBSomxM4 @s_channe pic.x.com/slGv0uNM6P
These channels provide health facilities with near real-time reporting, eliminated the barriers imposed by difficult access to technological resources in remote areas.
Until next time
After the presentations, the Epihack event was closed after an encouraging speech by Dr. Deo Mtasiwa from the Tanzanian government, expressing the willingness to follow up the work achieved during the event. The prototypes had the effect of ultimately concretising the discussions that had taken place throughout the week, and being able to see them operational caused an otherwise unachievable impact on the health experts, who could witness what could be created through technology by engaging developers in their field of work.
The Epihack was a fantastic event and a truly enriching experience, in which we had the possibility to interact with fellow developers from other parts of the world, observe first-hand the situation of communities in East Africa, and give a helping hand in solving the problem of disease detection.
Looking forward to the next one!
